The Evolutionary Influence of Ancient Retroviruses in Passeriform Genomes
- 演化之聲

- Mar 12
- 5 min read
Retroviruses are a group of viruses that reverse-transcribe their RNA genomes into DNA. The resulting DNA is inserted into the host cell's genome, after which the host's transcription and translation machinery produces viral RNA and proteins, allowing new viral particles to assemble and complete their replication cycle.

The genome of a retrovirus is flanked at both ends by long terminal repeats (LTRs). Through the action of the viral enzyme integrase, the viral genome becomes integrated into the host genome. Once inserted, it generally remains there permanently and becomes part of the host's genetic material.

If a retrovirus infects a germ cell and integrates into its genome, the viral sequence can be inherited by the next generation. Viral DNA that remains in the genome under these circumstances is known as an endogenous retrovirus (ERV). In most cases, the ERVs passed to descendants have already lost their ability to produce infectious viruses. Only fragments of viral genes or regulatory elements remain functional. Over many generations, cellular processes such as DNA recombination and ERV deletion further degrade these sequences, often leaving incomplete remnants.
Because many retroviruses infect only vertebrates, ERVs have played an important role in vertebrate evolution. In some cases they have beneficial effects, for example by becoming regulatory elements within host genomes. In other situations they have been linked to serious diseases. One example is human endogenous retrovirus K (HERV-K), which still retains an active envelope glycoprotein gene capable of producing viral proteins and has been associated with neurodegenerative diseases such as amyotrophic lateral sclerosis.
Host cells also possess mechanisms capable of removing ERVs. One such mechanism involves homologous recombination during DNA repair. In this process, recombination occurs between the two LTRs of an ERV, deleting the internal viral sequence. However, because the recombination occurs between the LTRs, one LTR remains in the genome afterward. This remaining fragment is known as a solo-LTR. As a result, ERVs in vertebrate genomes can exist in multiple forms: intact copies, incomplete fragments, or isolated solo-LTR remnants.

The distribution of ERVs varies across animal groups. Birds represent a particularly interesting case. Compared with mammals and reptiles, avian genomes contain a markedly lower proportion of ERVs, suggesting that birds possess distinctive genomic and evolutionary patterns.
A study published in 2024 examined ERVs across the genomes of 405 representative amniotes, including 362 bird species, 23 reptiles, and 20 mammals. The goal was to understand how ERVs have influenced avian evolution.
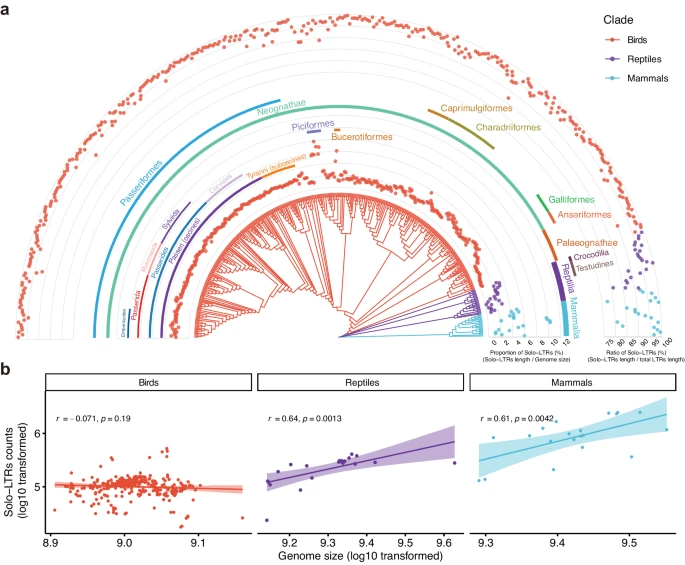
The analysis revealed that the proportion of ERVs in bird genomes is consistently lower than in other amniotes. At the same time, birds exhibit a higher frequency of solo-LTR formation. This pattern indicates that birds remove ERVs from their genomes more efficiently than many other vertebrate groups.
ERVs themselves can be divided into multiple categories based on their retroviral origins, including ERVK, ERVL, ERVW, and ERV1. When researchers examined the distribution of these ERV-derived solo-LTRs in bird genomes, they discovered a pattern that differed markedly in the order Passeriformes.

The earliest fossil record of Passeriformes dates to approximately 52 million years ago, with an estimated origin around 60 million years ago. This order contains at least 6,500 species and represents more than 60 percent of all bird species, making it the largest avian lineage. Because many of its members are skilled vocalizers, they are commonly referred to as songbirds.
The study found that most bird groups maintain a diverse set of ERV solo-LTRs. However, Passeriformes show a pronounced accumulation of ERVK solo-LTRs, which are derived from β-retroviruses (Betaretrovirus). The closer a lineage is to modern branches of Passeriformes, the greater the number of ERVK solo-LTRs it tends to possess. The highest accumulation occurs in the parvorder Passerida, which originated about 22.4 million years ago. Moreover, the accumulation of ERVK solo-LTRs shows a strong positive correlation with speciation events in Passeriformes, suggesting that diversification within this group was accompanied by the increasing presence of ERVK solo-LTRs.
To better understand the functional significance of ERVK solo-LTRs, researchers used the zebra finch (Taeniopygia guttata) as a model species. They discovered that up to 120 ERVK elements in the zebra finch genome still retain transcriptional activity. Their expression levels are particularly high in the ovary, testis, and primordial germ cells, significantly exceeding those observed in other tissues.
This pattern indicates that ERVK elements remain active in reproductive tissues. The RNA produced from ERVK can be reverse-transcribed again into DNA and inserted elsewhere in the genome, allowing the element to continue expanding. Subsequent deletion processes convert many of these insertions into solo-LTR fragments, gradually accumulating them in Passeriform genomes.
Researchers also compared gene expression patterns between the zebra finch and the chicken (Gallus gallus). They identified twenty genes in the zebra finch whose nearby genomic regions contain ERVK solo-LTR insertions that are absent in the chicken. These genes also exhibit significant differences in expression levels between the two species.
Among them, nine genes show higher expression in the zebra finch, while eleven display lower expression.
One particularly important example is the ITGA2 gene. In the zebra finch brain, this gene shows elevated expression in regions associated with vocal complexity. A fragment of ERVK solo-LTR sequence was found upstream of the gene, and experiments demonstrated that this element can enhance ITGA2 expression. This finding suggests that the insertion of ERVK solo-LTR near ITGA2 may have contributed to the evolution of vocal learning in Passeriform birds.
During the same geological period when ERVK expanded in Passeriform genomes—from the Paleogene into the Neogene—many other animal groups, including primates and rodents, also experienced increases in ERVK activity. However, ERVK elements in mammals often remain relatively intact and can still produce viral proteins, which has been associated with aging and cancer. In contrast, birds appear to convert ERVs into solo-LTR fragments more efficiently, reducing their harmful effects.
Even today, ERVK solo-LTR sequences continue to accumulate in Passeriform genomes.
In an unexpected twist of evolutionary history, an ancient viral fragment inserted upstream of the ITGA2 gene, enhancing vocal learning ability in these birds. Passeriformes are widely known for their diversity and complex vocal communication. In this case, genetic remnants from ancient viruses may have played a role in driving one of the most remarkable radiations in avian evolution.
Author: Bai Leng,Editor: Shui-Ye You
Reference:
Chen, G., Yu, D., Yang, Y., Li, X., Wang, X., Sun, D., Lu, Y., Ke, R., Zhang, G., Cui, J., Feng, S. (2024). Adaptive expansion of ERVK solo-LTRs is associated with Passeriformes speciation events. Nature Communications.




Comments